Preprocessing Data¶
The synomicsbench.processing module provides a comprehensive suite of utilities for cleaning, transforming, and integrating omics and clinical data. This processing pipeline minimizes the confounding impact of data quality on generative model effectiveness, ensuring unbiased comparability among the evaluated synthetic data generation methods. As illustrated in Figure 2, the pipeline follows five sequential steps: 1) Data Filtering, 2) Transcriptomic Harmonization, 3) Feature type classification, 4) Multivariate missing data imputation, and 5) Data Integration. The pipeline is flexible and configurable, allowing users to customize preprocessing steps based on their specific datasets and analysis requirements.
Data Integration Pipeline¶
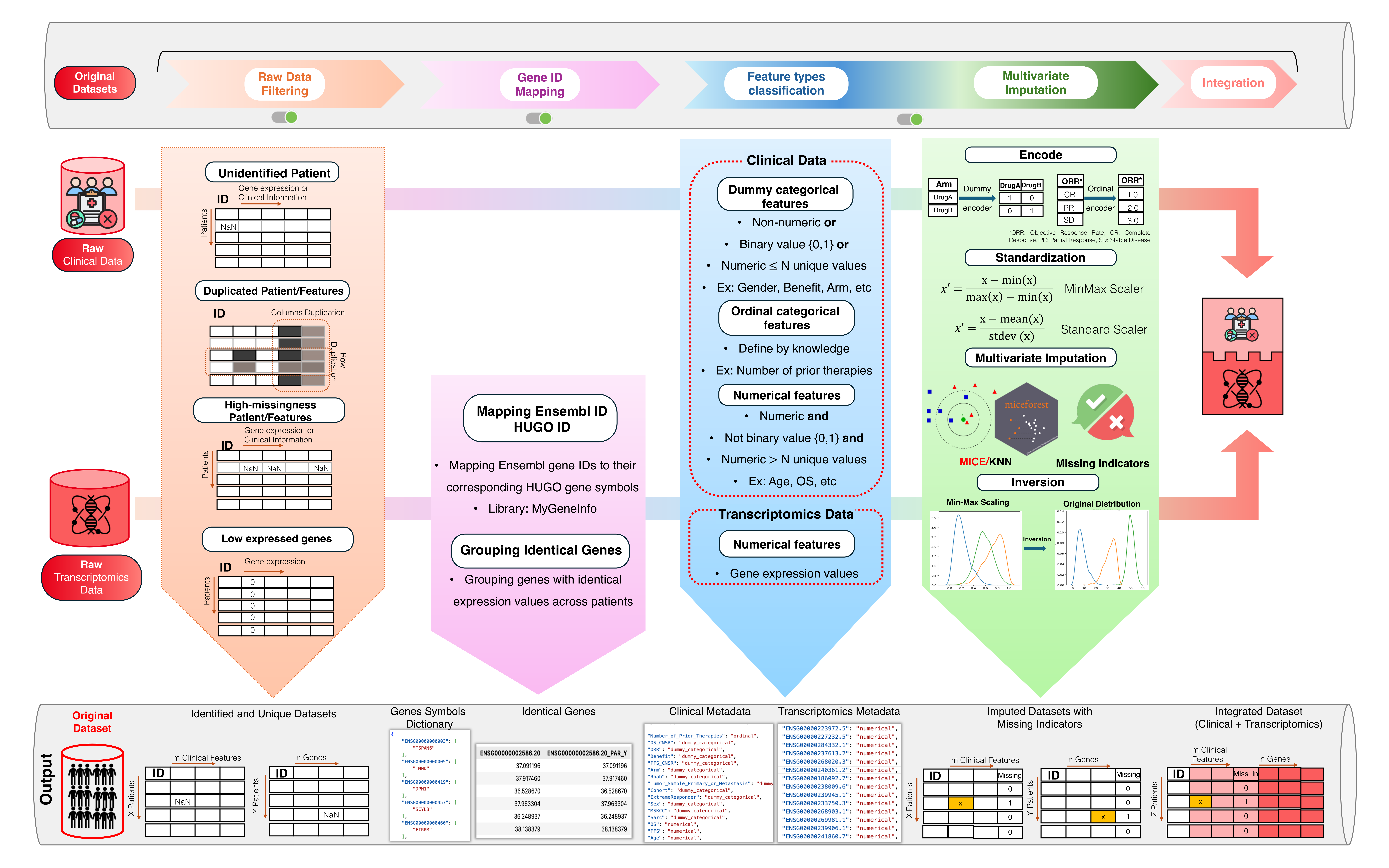
Figure 1: Standardized preprocessing pipeline for clinical-transcriptomic data integration.
The DataIntegrationPipeline ensures data quality and consistency before synthetic data generation or downstream analysis.
Pipeline Execution¶
The following example demonstrates how to configure and execute the complete data integration pipeline:
import pandas as pd
from synomicsbench.processing.pipeline import DataIntegrationPipeline
# Load your raw data
clinical_data = pd.read_csv("clinical_data.csv")
transcriptomics_data = pd.read_csv("transcriptomics_data.csv")
output_dir = "./integrationpipeline_output"
ordinal_cat_columns = [
"MSKCC",
"Number_of_Prior_Therapies",
"ORR",
"ExtremeResponder",
"Benefit",
]
steps_config = {
"remove_undefined": True,
"remove_duplicates": True,
"remove_overmissing_samples": True,
"remove_low_expression_genes": True,
"check_duplicate_genes": True,
"mapping_genes": True,
"feature_engineering": True,
"integrate_data": True,
}
imputer_params = {
"iterations": 10,
"n_estimators": 100,
"random_state": 42,
}
pipeline = DataIntegrationPipeline(
output_dir=output_dir,
logger="Integration_final",
)
results = pipeline.run_pipeline(
clinical_data=clinical_data,
transcriptomics_data=transcriptomics_data,
clinical_id_column="RNA_ID",
transcriptomics_id_column="Sample",
integration_id_column="Patient_ID",
steps_config=steps_config,
overmissing_samples_threshold=50,
overmissing_features_threshold=50,
unique_threshold=10,
scaler="minmax",
ordinal_cat_columns=ordinal_cat_columns,
imputer="mice",
imputer_params=imputer_params,
low_expression_variance_threshold=0.0005,
add_indicators=True,
verbose=True,
)
# Access outputs — run_pipeline() returns a dict with three keys
processed_clinical = results["processed_clinical"]
processed_transcriptomics = results["processed_transcriptomics"]
integrated_data = results["integrated_data"] # None if integrate_data=False
print(f"Clinical shape: {processed_clinical.shape}")
print(f"Transcriptomics shape: {processed_transcriptomics.shape}")
print(f"Integrated shape: {integrated_data.shape}")
The steps_config dictionary controls which preprocessing steps are executed, allowing flexible pipeline customization based on data characteristics and analysis requirements. The pipeline returns a dict with keys processed_clinical, processed_transcriptomics, and integrated_data (the inner-joined dataset, or None when integrate_data=False).
Pipeline Steps¶
The data integration pipeline executes configurable steps in sequence, controlled by steps_config.
remove_undefined → remove_duplicates → remove_overmissing_samples →
(transcriptomics only) remove_low_expression_genes → check_duplicate_genes → mapping_genes →
feature_engineering → integrate_data
| Step key | What it does | Where it runs | Implementation (code) | Key params |
|---|---|---|---|---|
remove_undefined |
Drop rows with missing IDs | Clinical + transcriptomics | DataProcessor.remove_unknown_entities(...) |
*_id_column |
remove_duplicates |
Remove exact duplicate rows/cols | Clinical + transcriptomics | DataProcessor.remove_duplications(..., axis=0/1) |
axis |
remove_overmissing_samples |
Drop rows with missingness > threshold | Clinical + transcriptomics | DataProcessor.remove_overmissing_entities(...) |
overmissing_samples_threshold |
remove_low_expression_genes |
Filter zero-sum / near-zero variance genes | Transcriptomics | DataProcessor.remove_low_expression_genes(...) |
low_expression_variance_threshold |
check_duplicate_genes |
Find identical expression profiles | Transcriptomics | GeneQuery.check_duplicates(...) |
(none) |
mapping_genes |
Map Ensembl IDs → HUGO symbols | Transcriptomics | GeneQuery.mapping_genes(...) |
(none) |
feature_engineering |
Type classification + encoding + scaling + imputation | Clinical + transcriptomics | DataProcessor.feature_engineering(...) |
data_type, imputer, imputer_params, scaler, unique_threshold, ordinal_cat_columns |
integrate_data |
Merge clinical + transcriptomics on common ID | Final integration | DataIntegrationPipeline.integrate_data(...) |
integration_id_column |
Tip
If you only need one step (e.g., just filtering, encoding, or imputation), you can call the preprocessing utilities directly—see the section below.
Utility Functions (DataProcessor)¶
All helper functions are available in synomicsbench.processing.preprocessing.DataProcessor. These are useful when you want to execute specific preprocessing steps without running the full DataIntegrationPipeline.
Available Methods¶
| Method | Description | Key Parameters |
|---|---|---|
remove_duplications |
Drops duplicate rows or columns | axis=0/1 |
remove_unknown_entities |
Removes rows with missing IDs | id_column |
remove_overmissing_entities |
Drops rows with missingness above threshold | threshold |
find_missing_percent |
Calculates missingness percentage | |
remove_overmissing_features |
Drops columns with missingness above threshold | threshold |
remove_low_expression_genes |
Filters near-zero variance genes | gene_id_column, variance_threshold |
encode_dummy_features |
One-hot encodes categorical data | |
encode_ordinal_features |
Ordinal encodes categorical data | |
standardization |
Scales numerical features | scaler='minmax'/'standard' |
mice_imputation |
Multiple Imputation by Chained Equations | iterations, n_estimators, add_indicators |
extract_missingindicator_columns |
Extracts generated missingness indicators | |
inverse_dummy_features |
Reverses dummy encoding | dummy_cat_columns |
inverse_ordinal_features |
Reverses ordinal encoding | encoder |
inverse_standardization |
Reverses scaling | scaler |
knn_imputation |
k-Nearest Neighbors imputation | n_neighbors, dummy_cat_columns, ordinal_cat_columns |
feature_engineering |
Type classification, encoding, scaling, and imputation pipeline | data_type, imputer, imputer_params, scaler |
Example Usage¶
The following example uses a realistic clinical-transcriptomic dataset (30 patients, 6 clinical features, 30 genes) to demonstrate each DataProcessor step individually.
import pandas as pd
import numpy as np
from synomicsbench.processing.preprocessing import DataProcessor
# ── Sample data ──────────────────────────────────────────────────────────────
N = 30
np.random.seed(42)
data = {
"Patient_ID": [f"P{i+1:02d}" for i in range(N)],
"Gender": np.random.choice(["M", "F"], N),
"Mstage": np.random.choice(["I", "II", "III", "IV"], N),
"Tx_Start_ECOG": np.random.choice([0, 1, 2], N).astype(float),
"numPriorTherapies":np.random.randint(0, 4, N).astype(float),
"biopsyContext": np.random.choice(["Primary", "Metastatic"], N),
"Age": np.random.randint(45, 80, N).astype(float),
}
for i in range(1, 31):
data[f"Gene_{i:02d}"] = np.random.normal(loc=5, scale=2, size=N)
df = pd.DataFrame(data)
# Introduce realistic missingness
df.loc[[10, 12], "Patient_ID"] = np.nan # undefined IDs → will be dropped
df.loc[2, "Age"] = np.nan
df.loc[5, "Mstage"] = np.nan
df.loc[15, "Tx_Start_ECOG"] = np.nan
# ── 1. Filtering & QC ────────────────────────────────────────────────────────
df_clean = DataProcessor.remove_unknown_entities(df, id_column="Patient_ID")
df_clean = DataProcessor.remove_duplications(df_clean, axis=0).reset_index(drop=True)
# ── 2. Encoding ──────────────────────────────────────────────────────────────
dummy_cat = ["Gender"]
ordinal_cols = ["Mstage", "Tx_Start_ECOG", "numPriorTherapies", "biopsyContext"]
cat_encoded = DataProcessor.encode_dummy_features(df_clean[dummy_cat])
ord_encoded, ord_encoder = DataProcessor.encode_ordinal_features(df_clean[ordinal_cols])
# ── 3. Scaling ───────────────────────────────────────────────────────────────
num_scaled, num_scaler = DataProcessor.standardization(df_clean[["Age"]], scaler="minmax")
# ── 4. MICE imputation ───────────────────────────────────────────────────────
# Operates on the cleaned DataFrame; missing indicators are appended automatically.
df_imputed = DataProcessor.mice_imputation(
df_clean.reset_index(drop=True),
iterations=10,
n_estimators=100,
add_indicators=True,
)
# ── 5. Extract missingness indicators ────────────────────────────────────────
indicators = DataProcessor.extract_missingindicator_columns(df_imputed)
print(indicators.columns.tolist())
# ['missingindicator_Age', 'missingindicator_Mstage', 'missingindicator_Tx_Start_ECOG']