Statistical Fidelity¶
Statistical fidelity evaluates how faithfully synthetic data reproduces the statistical properties of the original dataset at two levels: marginal distributions of individual features (univariate similarity) and pairwise relationships between features (bivariate similarity).
Univariate Similarity¶
Univariate similarity measures how well the distribution of each individual feature is preserved.
Metrics¶
- Numerical features — the Kolmogorov–Smirnov (KS) statistic measures the maximum distance between the cumulative distributions:
- Categorical features — the Total Variation Distance (TVD) quantifies the discrepancy between frequency distributions:
To ensure balanced evaluation across modalities, the final univariate score averages the mean scores of clinical and transcriptomic features separately:
where \(N\) and \(M\) denote the total number of clinical features and genes, respectively.
Code Example¶
from synomicsbench.metrics.fidelity.UnivariateSimilarity import UnivariateSimilarity
from synomicsbench.processing.metadata import MetaData
# Compute metadata (auto-detect feature types)
metadata = MetaData.get_metadata(
data=original_data,
threshold_unique_values=10,
ordinal_features=None
)
# Compute univariate similarity scores
uni = UnivariateSimilarity(output_dir="results/statistical_fidelity")
scores = uni.get_univariate_score(
original_data=original_data,
synthetic_data=synthetic_data,
metadata=metadata
)
detail_df = uni.get_detail_df()
print(f"Univariate Score: {scores:.4f}")
print(detail_df.head())
Bivariate Similarity¶
Bivariate similarity evaluates how well pairwise relationships between features are preserved.
Metrics¶
- Numerical–numerical pairs — Spearman's rank correlation (\(\rho\)):
- Categorical–categorical and mixed pairs — Cramér's V. For mixed pairs, numerical features are discretized into 10 bins:
The overall bivariate score equally weights three functional modalities:
where each \(\bar{S}_{\text{modality}}\) is the mean similarity score over all feature pairs within that category.
Code Example¶
from synomicsbench.metrics.fidelity.PairwiseSimilarity import PairwiseSimilarity
from synomicsbench.processing.metadata import MetaData
metadata = MetaData.get_metadata(
data=original_data,
threshold_unique_values=10,
ordinal_features=None
)
pairwise = PairwiseSimilarity(
original_data=original_data,
synthetic_data=synthetic_data,
metadata=metadata,
output_dir="results/statistical_fidelity",
name="my_dataset"
)
results = pairwise.get_pairwise_scores(method="spearman")
# results is a dict with keys: "PairwiseScore", "OriginalCorrelation", "SyntheticCorrelation"
scores = results["PairwiseScore"]
print(f"Mean Bivariate Score: {scores.mean():.4f}")
Visualization¶
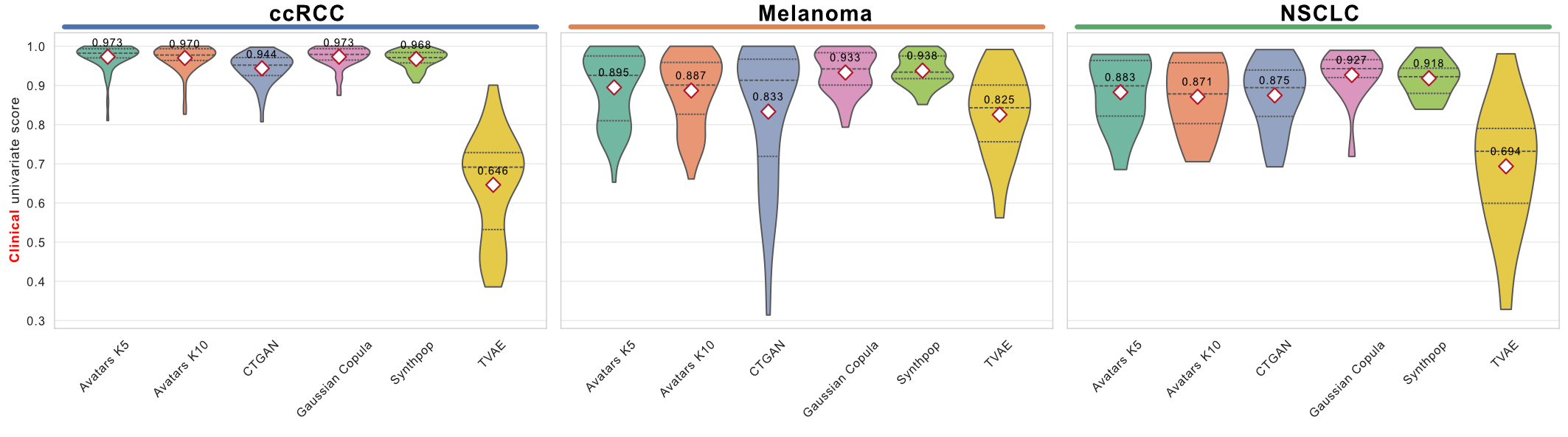
The violin plot below shows the distribution of per-feature similarity scores across SDG methods. The plot_violin_grid_by_cancer function produces manuscript-quality figures. The input dict must be structured as {cancer: {method: [scores]}}:
from synomicsbench.metrics.fidelity.visualization import plot_violin_grid_by_cancer
# cancer_to_method_scores: {cancer_name: {method_name: [score_replicate_1, ...]}}
# Each list contains per-feature (univariate) or per-pair (bivariate) scores across replicates.
Score_Dict = {
"ccRCC": {
"Avatars K5": avatars_k5_ccrcc_scores,
"Avatars K10": avatars_k10_ccrcc_scores,
"CTGAN": ctgan_ccrcc_scores,
"Gaussian Copula": gc_ccrcc_scores,
"Synthpop": synthpop_ccrcc_scores,
"TVAE": tvae_ccrcc_scores,
},
"Melanoma": {
"Avatars K5": avatars_k5_melanoma_scores,
# ...
},
}
fig, axes, mean_by_cancer = plot_violin_grid_by_cancer(
cancer_to_method_scores=Score_Dict,
value_name="Univariate Score",
figsize=(18, 5)
)