Survival Analysis¶
Survival analysis evaluates whether synthetic data preserves clinically meaningful survival patterns by comparing Overall Survival (OS) and Progression-Free Survival (PFS) between treatment responders and non-responders.
C-index Score¶
The C-index score measures similarity based on the absolute difference between original and synthetic concordance indices from Cox proportional hazards models:
\[C_\text{score}^\text{OS} = 1 - \left|C_\text{original}^\text{OS} - C_\text{synthetic}^\text{OS}\right|\]
\[C_\text{score}^\text{PFS} = 1 - \left|C_\text{original}^\text{PFS} - C_\text{synthetic}^\text{PFS}\right|\]
\[C_\text{score}^\text{final} = \frac{C_\text{score}^\text{OS} + C_\text{score}^\text{PFS}}{2}\]
Higher values indicate greater agreement in survival discrimination. Kaplan-Meier survival curves are compared using the log-rank test.
Code Example¶
from synomicsbench.metrics.narrow_utility.survival_analysis import SurvivalEvaluator
# Define dataset dictionary
survival_cols = ["OS", "OS_CNSR", "Benefit"]
original_data = pd.read_csv("original_data.csv")
synthetic_data = pd.read_csv("synthetic_data.csv")
dataset_dict = {
"Origin": original_data[survival_cols],
"Synthetic_data": synthetic_data[survival_cols],
}
# Define phenotype comparison
phenotype = {"Benefit": ["CB", "NCB"]}
# Define evaluator
evaluator = SurvivalEvaluator(
datasets_dict=dataset_dict,
phenotype=phenotype,
time_target="OS",
event_target="OS_CNSR",
original_name="Origin"
)
# Compute C-index similarity scores with respect to the original data
summary_df = evaluator.compute_survival_metrics()
scored_df = evaluator.compute_cindex_scores(original_name="Origin")
# Extract the C-index score
synthetic_score = scored_df.loc[scored_df["Dataset"] == "Synthetic_data", "C-index_score"].values[0]
print(f"Synthetic Score: {synthetic_score:.4f}")
Kaplan-Meier Curve Visualization¶
from synomicsbench.metrics.narrow_utility.survival_analysis import SurvivalEvaluator
# Define datasets — the key for the reference must match original_name
datasets = {
"Origin": real_df,
"Gaussian Copula": gc_df,
"Avatars K5": avatars_k5_df,
"Avatars K10": avatars_k10_df,
"CTGAN": ctgan_df,
"Synthpop": synthpop_df,
"TVAE": tvae_df,
}
phenotype = {"Benefit": ["CB", "NCB"]}
evaluator = SurvivalEvaluator(
datasets_dict=datasets,
phenotype=phenotype,
time_target="OS",
event_target="OS_CNSR",
original_name="Origin"
)
evaluator.compute_survival_metrics()
fig, summary = evaluator.plot_grid(
save_dir="survival_analysis_results/"
)
# Each panel shows: Kaplan-Meier curves, log-rank P value, C-index
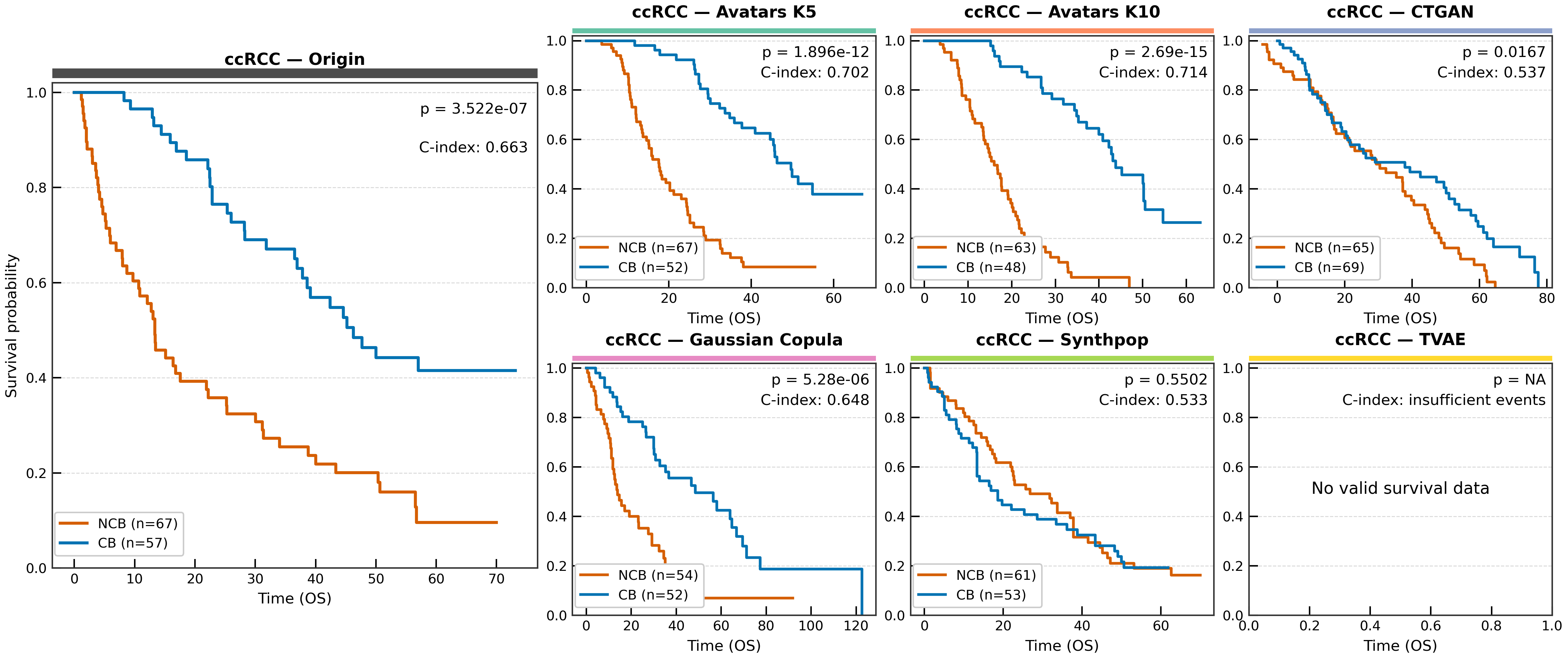
Survival signal preservation across SDG methods. Log-rank P values and C-indices are annotated in each panel.